About Us
We envision a world where all Oakland children read at or above grade level, no one suffers from Inflammatory Bowel Disease and Bay Area artists thrive.
Who We Are
Meet the Board members and staff who are at the core of our work.
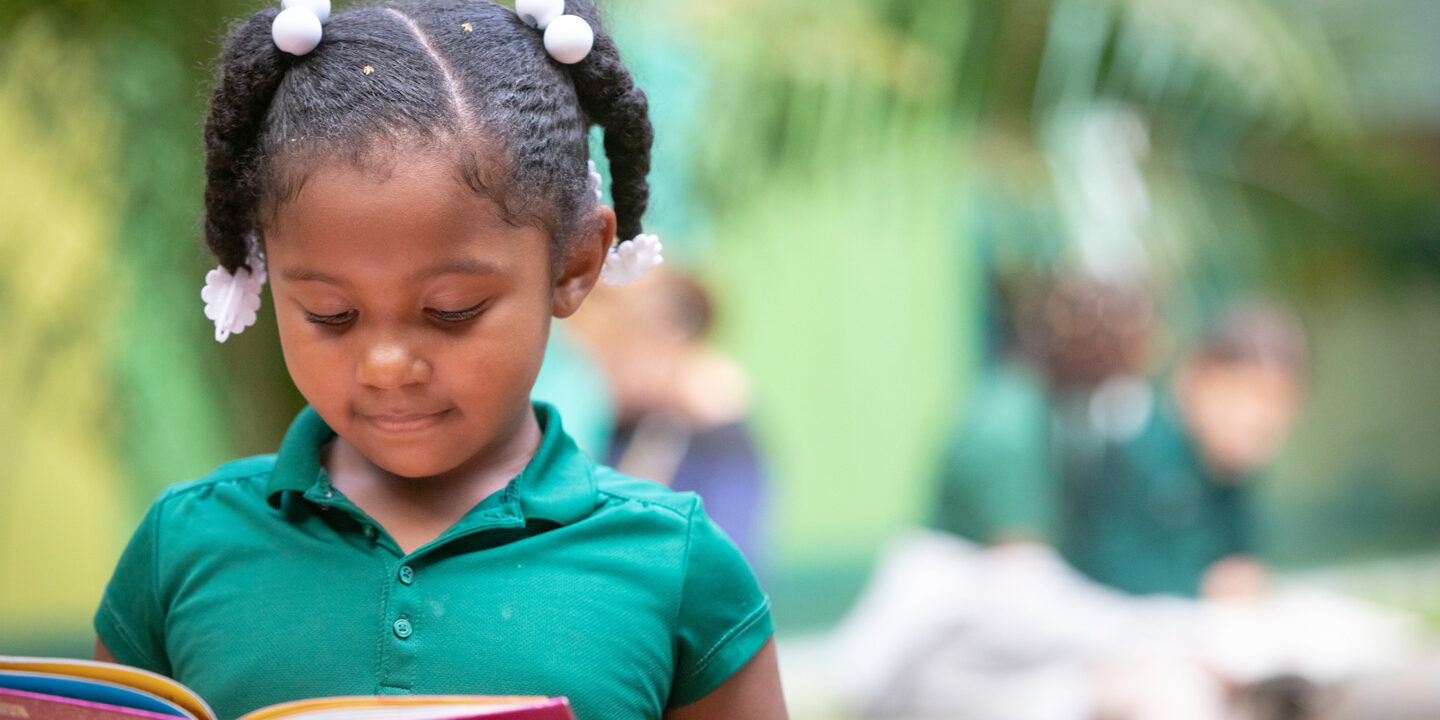
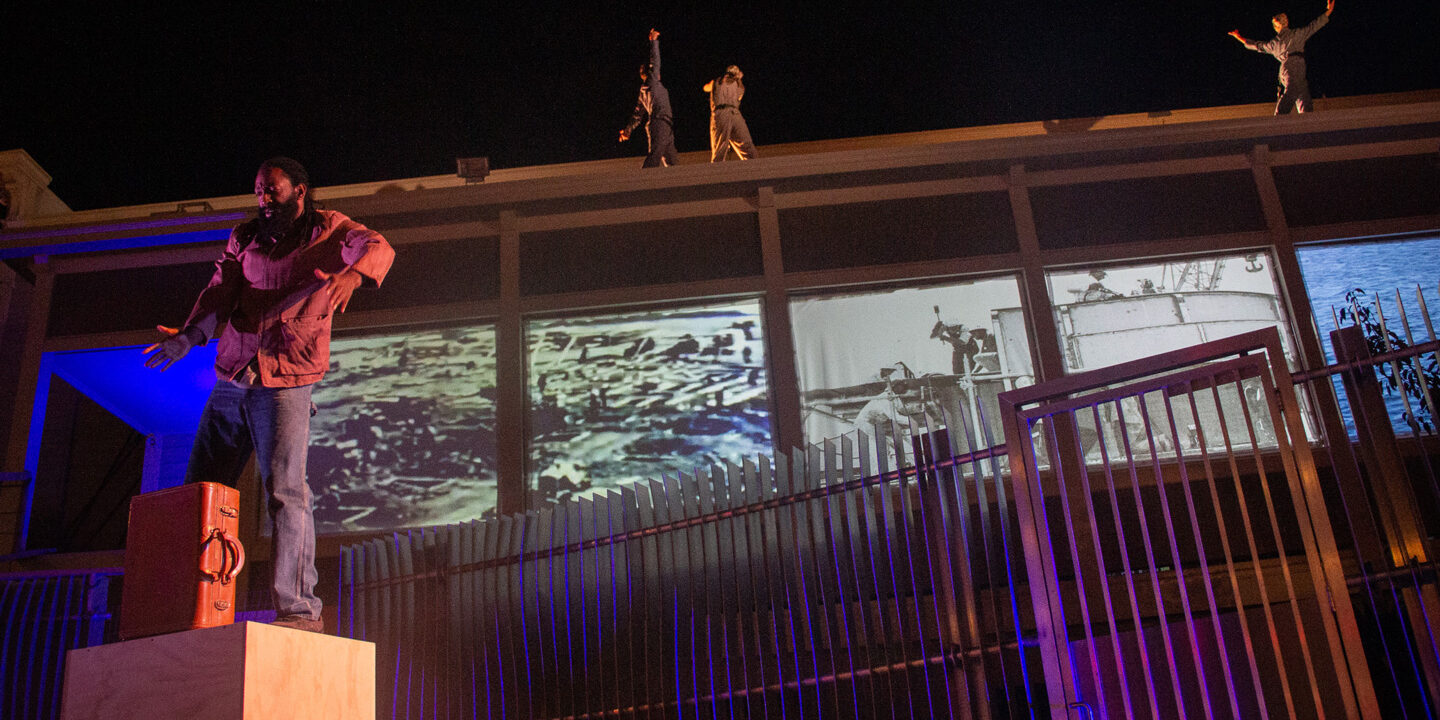
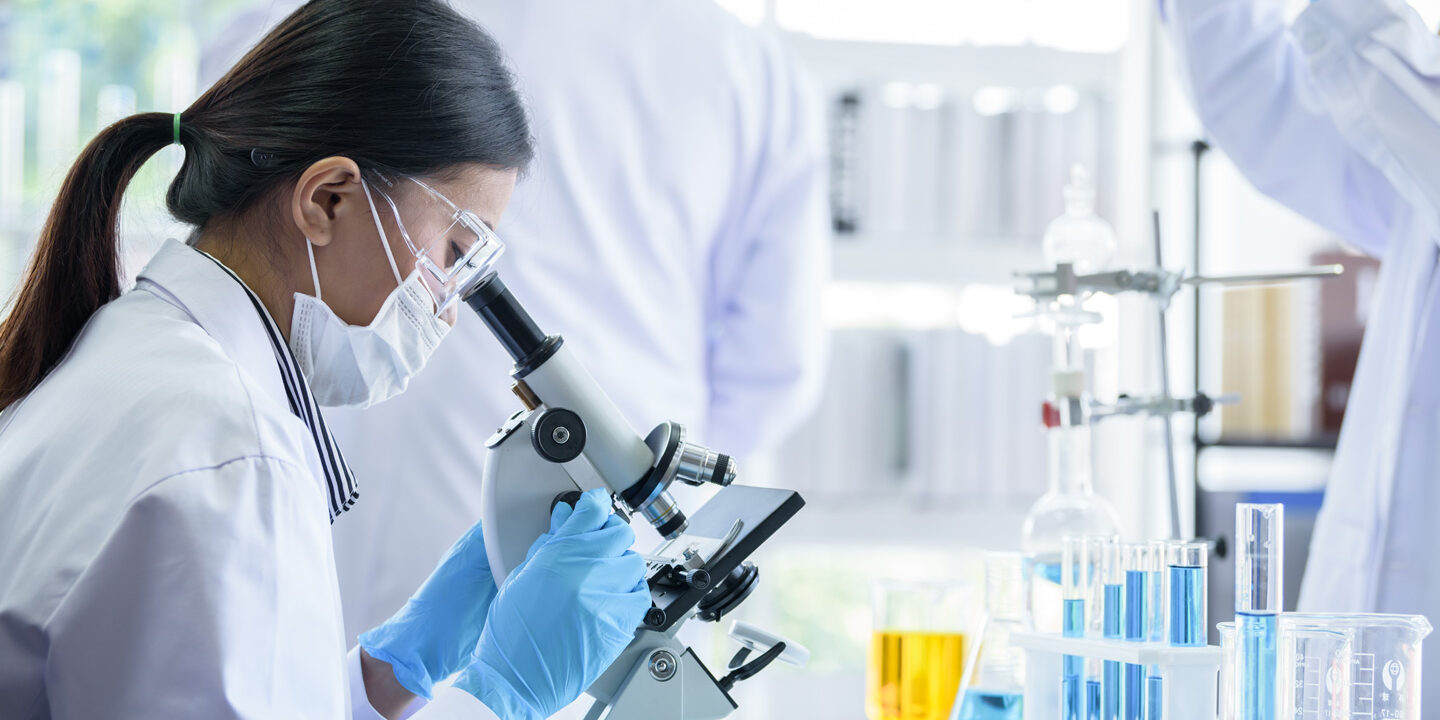
What We Fund
Learn more about our grantmaking in the Arts, Education and Health.
What We’re Learning
Read research and reports that share what we’re learning from our work with the communities we serve.
Grants
Collaboration and innovation are at the heart of all our programs. Read more about our grant opportunities.
Foundation News & Stories
Read the latest news from the Foundation and learn about the organizations we support.
Fifteen Years Of Lessons At The Kenneth Rainin Foundation
02/14/2024 | Foundation News
As we mark our 15th anniversary, CEO Jen Rainin shares valuable lessons learned along the way and how they have shaped the Foundation.
04/24/2024 | Arts
Meet the 2024 Rainin Fellows—Adrian L. Burrell (Film), Antoine Hunter, Purple Fire Crow (Dance), Ayodele “WordSlanger” Nzinga (Theater), and TNT Traysikel (Public Space)
01/23/2024 | Education
2023 end-of-year Education grants support Oakland’s early learning ecosystem.
04/09/2024 | Health
Rainin Foundation Scientific Advisory Board Member Dr. David Artis’ groundbreaking research has been influential to understanding IBD. In this interview he talks about gut health and disease along with emerging opportunities to support innovation.
Sign Up For Our Newsletter
Stay updated on the Kenneth Rainin Foundation and our grantees.